Emergence comprises some 100 experts, serving as speakers, advisors or mentors, that guide how to identify societal needs and carry out the entrepreneurial process.
Category: Bioengineering
Stanford Biodesign fellows hope to spur innovation in home countries
Biodesign program trains global fellows to take what they learn about technological innovation back home to train others.
Catalyst’s newest cohort spotlights Stanford innovation
Stanford Medicine's Catalyst program, which aims to accelerate impactful health care innovations, launches a new cohort.
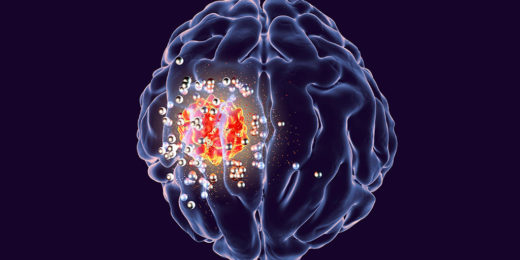
Wireless implant could help remove deadly brain tumors
Brain tumors are among the most deadly and difficult-to-treat cancers. Glioblastoma, a particularly aggressive form, kills more than 10,000 Americans a year and has a …
Unconventional Paths: Sneaky submarines and super surgeries
Bioengineer Alison Marsden uses computer modeling skills honed on submarines to help surgeons plan the best repairs for babies' hearts.
Engineering a new heart, layer by layer
Stanford researchers are building a heart through tissue engineering techniques in the hopes of better treating congenital heart defects.
‘Haptic emojis’ can convey emotions through simulated touch
Scientists have devised a wearable sleeve that helps communicate physical touch from afar, a concept dubbed "haptic communication."
Saving the world with synthetic biology
A Stanford Medicine bioengineer sets out to create a world fueled by synthetic biology, creating tools and technologies to see it through.
When it comes to healing without scarring, it pays to be small
Researchers have found that a phenomenon tied to animal size helps determine whether animals heal without scarring after burn injury.
Stanford Medicine magazine: Unlocking the brain’s mysteries
This new issue of Stanford Medicine magazine explores scientific advances that are helping unlock the mysteries of the brain.
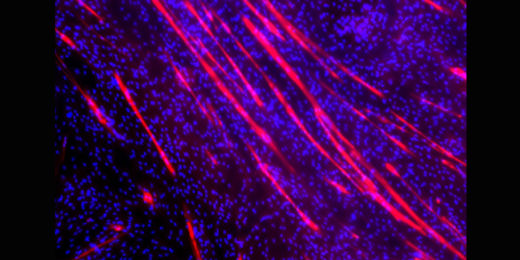
Engineered tissue sent into space to test muscle loss drugs
To help us understand muscle loss as we age, a Stanford Medicine research team’s engineered tissue is sent to the International Space Station.
Pain relief device uses real-world evidence to gain clearance, expanding options for kids
Tracking a pain-relief device's success in patients who aren't in clinical trials is seen as a promising approach to expanding treatment options for kids.
Easier-to-use technology helps young people with type 1 diabetes
Technology that sends blood sugar-level updates to their smartphones improves outcomes for young people with type 1 diabetes, a Stanford trial shows.
How does 2020 Nobel Prize-winning CRISPR technology work?
The 2020 Nobel Prize in Chemistry recognized the scientists who developed the CRISPR-Cas9 gene-editing technology. Here's how it's changing medicine.
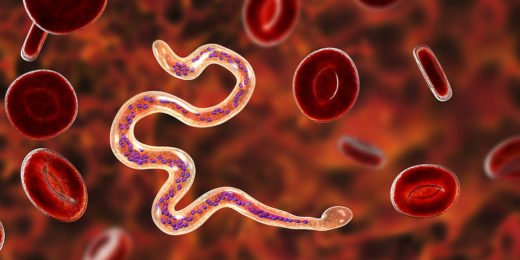
Stanford students design a device to detect early-stage river blindness
A team of Stanford undergraduates designed a device that uses blue-light imaging technology to diagnose a parasitic disease called river blindness.
AI researchers explore solutions for real-life health challenges
A device to prevent falls and another to better diagnose people with developmental disorders are among the AI projects funded under a new grant program.